Denaturation and Renaturation is the characteristic of DNA molecules. The factors like temperature, pH and chemical agents form or break the hydrogen bonds present between the two strands of DNA. It is a reversible process and it is used to study the complexity of the genome. Genetic Research & such studies use DNA Denaturation and Renaturation processes.
Denaturation is defined as change in the native structure of the biomolecule. Breaking of non covalent bonds like hydrogen, ionic and van der waals forces etc causes the denaturation. The DNA is right handed helical structure, it consist of two polynucleotide that are hold together by hydrogen bonds that are formed between the complementary nitrogen base pairs. DNA denaturation means the breaking of hydrogen bonds that causes separation of two strands.
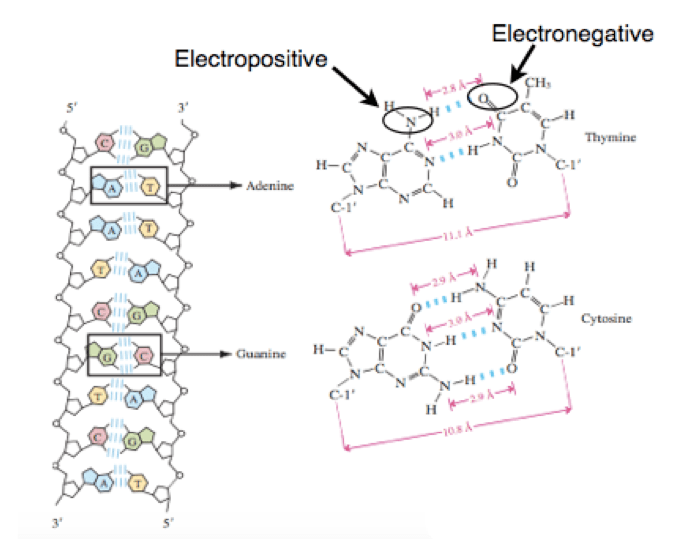
The hydrogen bond is the force of attraction between hydrogen atom of one covalently electronegative atom or group with other electronegative atom or group. It is weak bond and its strength ranges from 4 kJ to 50 kJ per mole. The strength of hydrogen bond is only 5% as compared to covalent bond. Due to the weakness, hydrogen bonds are prone to break. Different factors like temperature, pH and chemical agents can cause the breaking of hydrogen bonds. And hence, denaturation is also affected by the same factors.
- Temperature:
The temperature more than 80°C causes DNA denaturation by breaking the hydrogen bonds and producing two separate strands. The temperature at which DNA denature is called as melting point of DNA or melting temperature (Tm). The melting point is characteristic of DNA because Tm is affected by ATGC content. The sequence with more AT content would have less Tm then the DNA sequence with GC. It is because GC has triple bonds and hence more temperature or energy is required to break three bonds than two.
- pH:
The increase or decrease pH causes changes in ionic charges of nitrogen bases (purine and pyrimidines). At low pH, the amino groups become protonated and high pH causes its deprotonation. Both the conditions are not favorable to form the hydrogen bonds and hence it disrupts the native structure of DNA and lead to denaturation.
- Chemical agents:
Urea and formamide or formaldehyde is known to break the hydrogen bonds of DNA helix. These chemicals tend to form hydrogen bond with the electronegative centers of nitrogen bases. This causes disruption of the native hydrogen bonds between complementary base pairs of DNA structure. The melting point of DNA or Tm can be altered by adding such chemicals.
Characteristics of Denatured DNA
1) Viscosity: The double stranded helical structure is viscous in natured. When DNA is denatured by any of the mentioned factor, it loosed it viscosity.
2) Optical rotation: The native structure of DNA is optically activity. The denaturation of DNA causes decrease in the optical activity.
4) UV absorption: The native structure of DNA absorbs UV light at 260nm because of the aromatic rings of nitrogen bases. When DNA denatures the absorbance of UV light increase because of the exposure of Nitrogen bases and it is called as hyperchromic shift or hychromacity.
Hence the extent of denaturation of DNA can be observed of study by the UV light absorption.
PS: Denaturation and Renaturation of DNA is also used to understand the relativity of two genomes and repetitive sequences present in a genome.
Renaturation:
When the two separated strands of DNA bind together again, it is called as renaturation of DNA. It means it is transformation from denatured structure to native one. Renaturation of DNA occurs at room temperature or neutral pH or in the absence of denaturating chemical agents. The low temperature reduces the entropy and favors the colliding and contact of the two strands. When two strands come in to close proximity, they tend to form hydrogen bonds again forming double stranded helix. Hence Denaturation is reversible process.
When denatured DNA is renatured, the absorbance of UV at 260 nm decreases because of concealing of nitrogen bases. And this is called as Hypochromic shift.
The Single stranded DNA also tends to form hydrogen bonds with SS DNA from different source. It happens when SS DNA from different source have complementary sequence. This process is called as hybridization. It is used to study the evolutionary relationship of different species.
The renaturation of depends on concentration of DNA and time. Higher the concentration of DNA, faster is the renaturation. And hence the term is given as Cot (concentration x time)
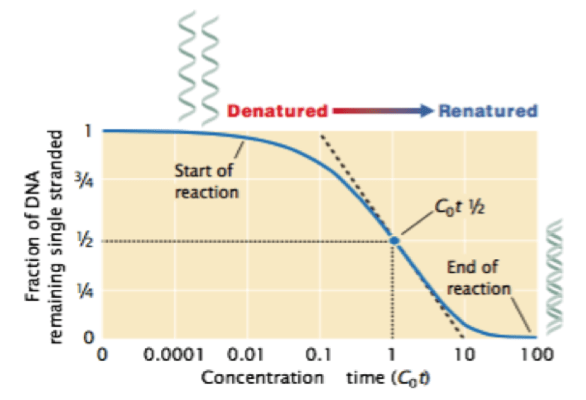
The plot depicts the renaturation of double stranded DNA of prokaryotic cell. The cot (concentration of DNA x time) is on X-axis and Percentage or fraction of SS DNA is on Y-axis. The curve that starts from upper left hand of Y-axis indicates the starting of renaturation reaction. 1 or 100% on X-axis means that the entire DNA is present in single stranded form. As the time passes, the DNA strands reanneal and the proportion or percentage of single stranded DNA decreases. When the entire DNA gets reannealed or renatured, the proportion of SS DNA become zero. The point at which half of the DNA is renatured, it is called as Cot1/2.
DNA contains unique and repetitive sequences. Unique means genes for particular character, which are present only once, or twice in DNA. Repetitive sequence means a stretch of nucleotide that is repeated multiple times in DNA. Britten and Davidson have found that higher the number of repetitive sequence present in DNA faster is the renaturation or annealing process. As the same sequence is in multiple copies, the probability of colliding with its complementary sequence becomes more. The repetitive sequences could be highly or moderately repeated.
To understand above statement, lets take an example. Imagine a tub with red, blue and white balls. Suppose, the box has 60 Red balls, 30 blue balls and 10 white balls. If you randomly try to pick these balls the probability of picking red balls is more than blue balls and much more than white balls. In the similar way, the collision of repetitive sequence with each other is more than unique sequence.
Hence, based on types of sequence (highly repetitive, moderately repetitive or unique genes) present in DNA affect the rate of renaturation. Rate of renaturation is directly proportional to number of repetitive sequences. The Cot analysis is used to measure repetitive sequences and complexity of genome.
The denaturation and renaturation of DNA plays an integral role in many methods and applications and are considered as an principal and crucial step in practical applications.
Reference:
https://www.chem.purdue.edu/gchelp/liquids/hbond.html
https://faculty.tru.ca/dnelson/courses/biol335/335notes/3recdna/4-hybrization/recDNA3a.html
D. Nelson and M, Cox, Freeman, 2004,
Biochemistry (9th Edition) Jeremy M. Berg, Lubert Stryer, John Tymoczko, Gregory Gatto, WH Freeman
If you liked this resource, please Like, Share, and Subscribe us for more content.



One thought on “Denaturation and Renaturation of DNA”