DNA replication is the process of DNA synthesis.
DNA is a double helical structure. It is a genetic material. It stores the information for protein synthesis. In Eukaryotes, it is present in nucleus and in prokaryotes it is present in cytoplasm in the form of nucleoid. When cell multiplies, the genetic material is transferred from one generation to another .
To transfer the genetic material to next generation, the cell needs to synthesize its replica. The replication of DNA allows to transfer its identical copy to its daughter cells. The DNA replication is called as semi-conservative because after replication out of two strands of DNA, one is original and other is newly synthesized. Hence, both the daughter cells receives one parental and one newly synthesized strands.
The process of synthesizing new DNA is called as replication. In eukaryotes, this process occurs in the S phase also called synthetic phase. The gap phase prepares all the essential factors that are required for replication. The S phase is followed by M phase, where the replicated DNA gets separated. Once the replicated DNA is separated, cytokinesis occurs and daughter cells are formed.
The replication process is of three steps Initiation, Elongation and Termination. The long length of DNA makes this process complex.
Initiation step of DNA replication:
The replication is initiated at a specific point and it is called as origin. The Origin is recognized and identified by the replication initiator protein.
In most of the prokaryotes, the cell has single origin. In E. coli the origin is 240 bp long. It is AT rich and hence contains double hydrogen bond between the complementary Adenine and Thymine. As AT contains only two bonds between them as compared to GC. It makes easy to break the bonds between AT and separate the strands. In eukaryotes, the DNA has multiple origins.
The Replication initiator protein recruits other essential proteins and enzymes for initiating the replication. When all the required proteins and enzymes are employed at origin it forms replication initiation complex. It is also referred as replisome.
DNA is double helical structure. The two strands are inter-wined to each other. Both the strands are hold together by hydrogen bonds. The hydrogen bonds are formed between complementary base pairing. For replication, the two inter-wined strands need to be separated. Using ATP, Helicase enzyme breaks the hydrogen bonds and unwinds the DNA.
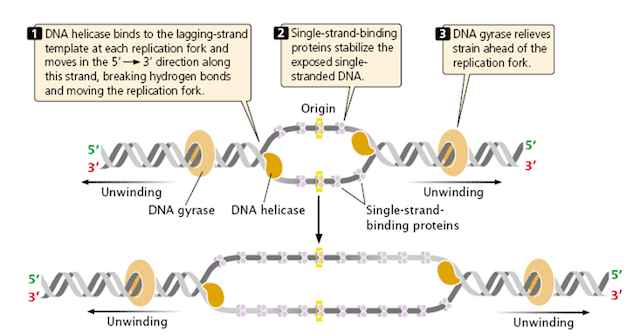
Replication fork
The unwinding of strands causes the formation of Y shaped replication fork. As both the strands unwinds, the replication forks are formed in both the direction forming replication bubble. During the replication, the bubble grows in both the direction.
When the strands are separated, they become unstable. Single stranded binding proteins are employed to stabilize the separated strands. Unwinding of strands, allows the recruitment of Primase for both the strands. It synthesize a small RNA strand called as RNA primer. It is complementary to the template and it provides its 3′ end for adding free nucleotides. Generally, the primer size ranges from 5 to 10bp. The RNA primer start off the strand synthesis by recruiting DNA polymerase at both the ends of replication fork (replication bubble).
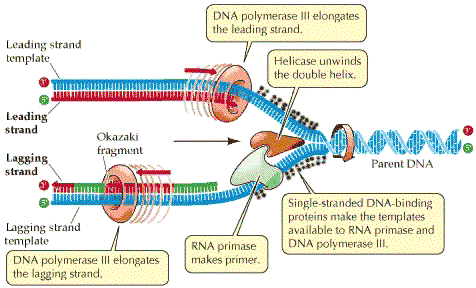
Elongation of DNA:
After binding of Polymerase to unwinded original strands (also called as template strand) synthesis of new strand is initiated. It consist of two antiparallel strands. One strand runs in 5′ to 3′ direction whereas the other one runs in the opposite direction. Both the opposite strands needs to be replicated.
Leading and Lagging strand:
The DNA polymerase can add free nucleotides at 3′-OH of RNA primer and extend the strand synthesis. The polymerase moves from 3′ to 5′ direction and synthesizes new strand continuously in 5′ to 3′ direction. The continuously growing strand is called as leading strand.
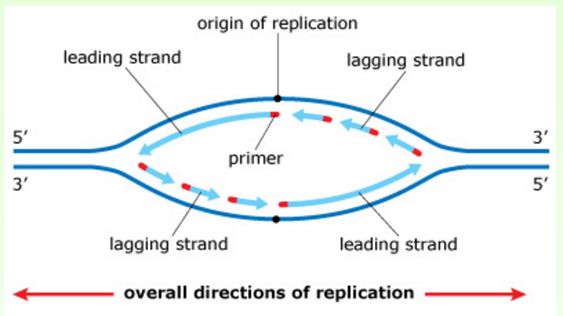
However, it is not possible to synthesize other strand continuously because of the Polymerase limitation. Polymerase cannot add free nucleotide at 5′ end. Due to which the other strand is synthesized in short fragments called as Okazaki fragments. The short fragments are sealed and linked with each other by ligase enzyme. As the rate of okazaki fragments synthesis is slower than continuous strand synthesis, it is called as lagging strand.
Sliding clamp and DNA topoisomerase are called as the maintenance protein. The sliding clamp holds the DNA polymerase at its right position. The topoisomerase plays an important role in relaxing DNA. When Helicase unwinds the strands and it causes topological strain which may be lead to damage the strand. Topoisomerase relives the strand by making temporary nicks and sealing it again and maintaining the DNA topology.
The DNA polymerase I is also a maintenance enzyme. It has exonuclease activity, which enable it to removes the short RNA primer. and refill the gap with the help of DNA ligase.
Termination of DNA replication:
The DNA replication terminates when the whole template is copied. In case of eukaryotes, as there are multiple origins, multiple replication forks are formed. When these forks meet, it eventually leads to terminate the replication process. The set of proteins and enzymes involved in termination process are different in prokaryotes and eukaryotes.
References:
https://teachmephysiology.com/biochemistry/cell-growth-death/
https://www.khanacademy.org/science/ap-biology/gene-expression-and-regulation/replication/a
Microbiology by Prescott, 5th Edition/
Dr. Sangha Bijekar has 9 years of Teaching Experience at University level. She loves to get engage in teaching and learning process. She is into blogging from last two years. She intends to provide student friendly reading material. She is avid Dog Lover and animal rescuer. She is learned Bharatnatyam and Katthak Dancer. She is into biking and She also loves to cook.


2 thoughts on “DNA Replication Process – A quick Read”